scvelo.tl.rank_velocity_genes¶
-
scvelo.tl.rank_velocity_genes(data, vkey='velocity', n_genes=100, groupby=None, match_with=None, resolution=None, min_counts=None, min_r2=None, min_corr=None, min_dispersion=None, min_likelihood=None, copy=False)¶ Rank genes for velocity characterizing groups.
This applies a differential expression test (Welch t-test with overestimated variance to be conservative) on velocity expression, to find genes in a cluster that show dynamics that is transcriptionally regulated differently compared to all other clusters (e.g. induction in that cluster and homeostasis in remaining population). If no clusters are given, it priorly computes velocity clusters by applying louvain modularity on velocity expression.
scv.tl.rank_velocity_genes(adata, groupby="clusters") scv.pl.scatter(adata, basis=adata.uns["rank_velocity_genes"]["names"]["Beta"][:3]) pd.DataFrame(adata.uns["rank_velocity_genes"]["names"]).head()
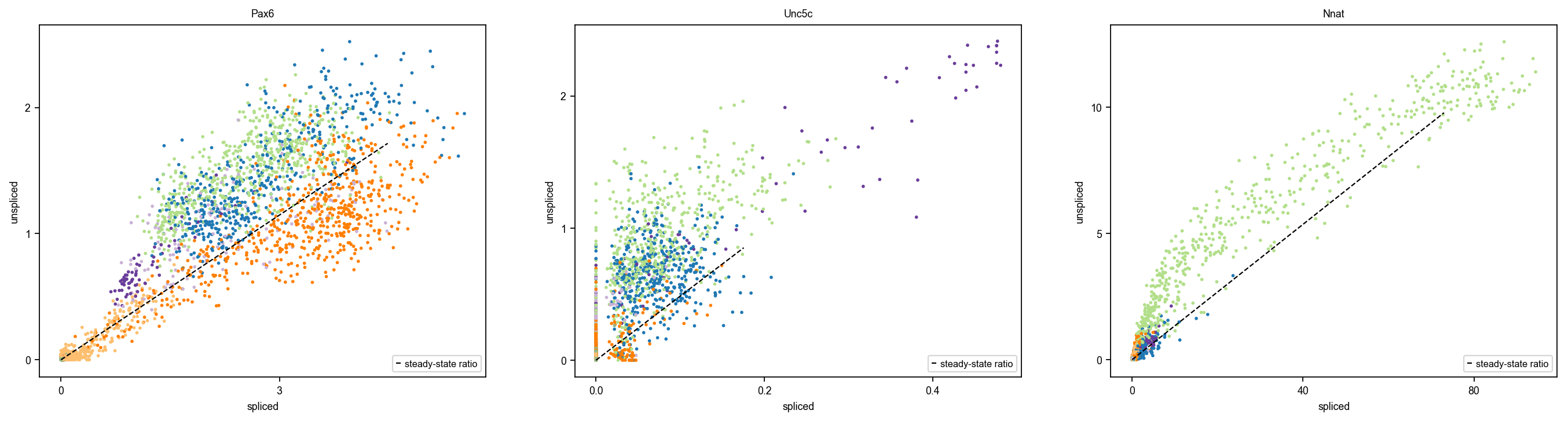

- data
AnnData Annotated data matrix.
- vkey: str (default: ‘velocity’)
Key of velocities computed in tl.velocity
- n_genesint, optional (default: 100)
The number of genes that appear in the returned tables.
- groupby: str, list or np.ndarray (default: None)
Key of observations grouping to consider.
- match_with: str or None (default: None)
adata.obs key to separatively rank velocities on.
- resolution: str or None (default: None)
Resolution for louvain modularity.
- min_counts: float (default: None)
Minimum count of genes for consideration.
- min_r2: float (default: None)
Minimum r2 value of genes for consideration.
- min_corr: float (default: None)
Minimum Spearmans correlation coefficient between spliced and unspliced.
- min_dispersion: float (default: None)
Minimum dispersion norm value of genes for consideration.
- min_likelihood: float between 0 and 1 or None (default: None)
Only rank velocity of genes with a likelihood higher than min_likelihood.
- copy: bool (default: False)
Return a copy instead of writing to data.
- Returns
rank_velocity_genes (.uns) – Structured array to be indexed by group id storing the gene names. Ordered according to scores.
velocity_score (.var) – Storing the score for each gene for each group. Ordered according to scores.
- data